Welcome to Nanocompore documentation
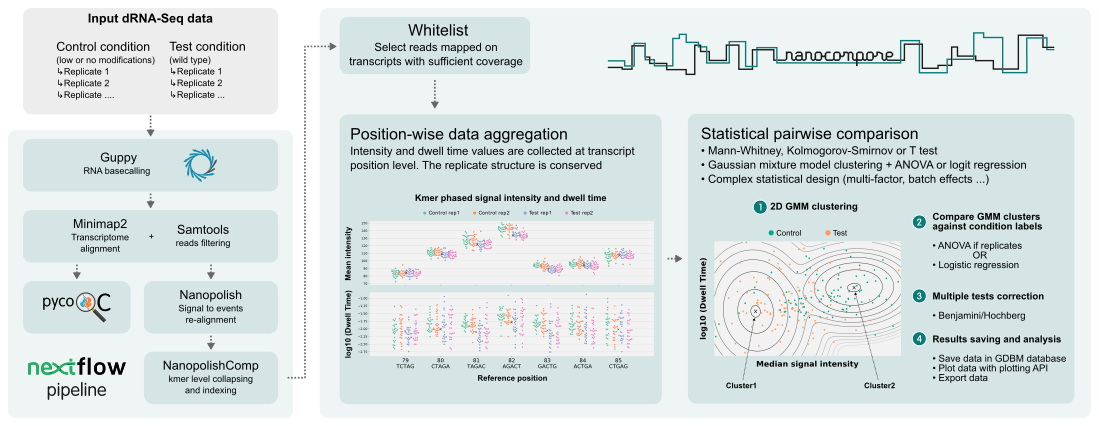
Nanocompore identifies differences in ONT nanopore sequencing raw signal corresponding to RNA modifications by comparing 2 samples
Nanocompore compares 2 ONT nanopore direct RNA sequencing datasets from different experimental conditions expected to have a significant impact on RNA modifications. It is recommended to have at least 2 replicates per condition. For example one can use a control condition with a significantly reduced number of modifications such as a cell line for which a modification writing enzyme was knocked-down or knocked-out. Alternatively, on a smaller scale transcripts of interests could be synthesized in-vitro.
Companion repositories
- NanoCompore_pipeline: Nextflow pipeline to preprocess data for NanoCompore
- Nanocompore_analysis: Analyses performed with Nanocompore for the BioRxiv preprint
- NanopolishComp: Collapse Nanopolish eventalign output per kmer, required before running NanoCompore